Method
A small VGG with a regression head.
The architecture is a sequential convolutional network with four
progressively deeper blocks. Each block holds two or three 3×3
convolutions, max-pooled and dropped at 0.3. The dense
head regresses a single non-negative scalar — the predicted cell
count for that 128×128 tile.
A ZeroPadding2D(40, 40) after the first convolution
expands the receptive field before the deeper layers compress it.
The full image is split into a uniform grid of approximately
100 px tiles, each resized to 128×128×3, batched through the
network. The total cell count is the ceiling of the summed
predictions.
Training: MSE loss under Adam at lr = 1e-3,
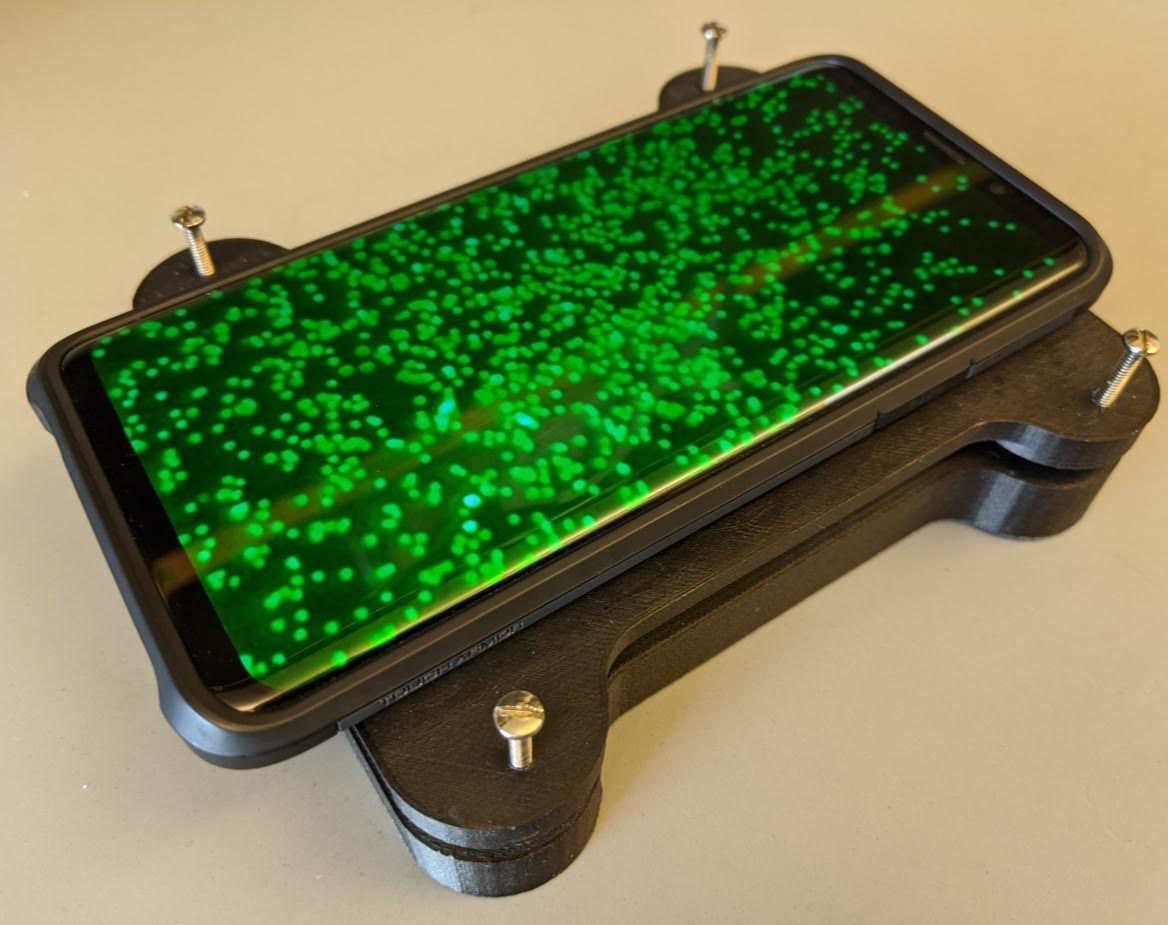
batch size 16, on fluorescent micrographs of human leukocytes and
spherical particles. Architecture, dataset curation, and training
pipeline by Harshitha Govindaraju (M.S. thesis,
Rutgers, 2021).